Introduction¶
NOvel Materials Discorvery (NOMAD) comprises storage, processing, management, discovery, and analytics of computational material science data from over 40 community codes. The original NOMAD software, developed by the NOMAD-coe project, is used to host over 50 million total energy calculations in a single central infrastructure instances that offers a variety of services (repository, archive, encyclopedia, analytics, visualization).
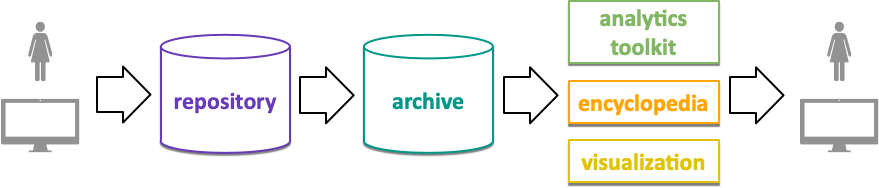
This is the documentation of nomad@FAIRDI, the Open-Source continuation of the original NOMAD-coe software that reconciles the original code base, integrate it’s services, allows 3rd parties to run individual and federated instance of the nomad infrastructure, provides nomad to other material science domains, and applies the FAIRDI principles as proliferated by the FAIRDI Data Infrastructure e.V.. A central and publically available instance of the nomad software is run at the MPCDF in Garching, Germany. Software development and the operation of NOMAD is done by the NOMAD Laboratory
The nomad software runs SAAS on a server and is used via web-based GUI and ReSTful API. Originally developed and hosted as individual services, nomad@FAIRDI provides all services behind one GUI and API into a single coherent, integrated, and modular software project.
This documentation is only about the nomad software; it is about architecture, how to contribute, code reference, engineering and operation of nomad.
Architecture¶
The following depicts the nomad@FAIRDI architecture with respect to software components in terms of python modules, gui components, and 3rd party services (e.g. databases, search engines, etc.). It comprises a revised version of the repository and archive.
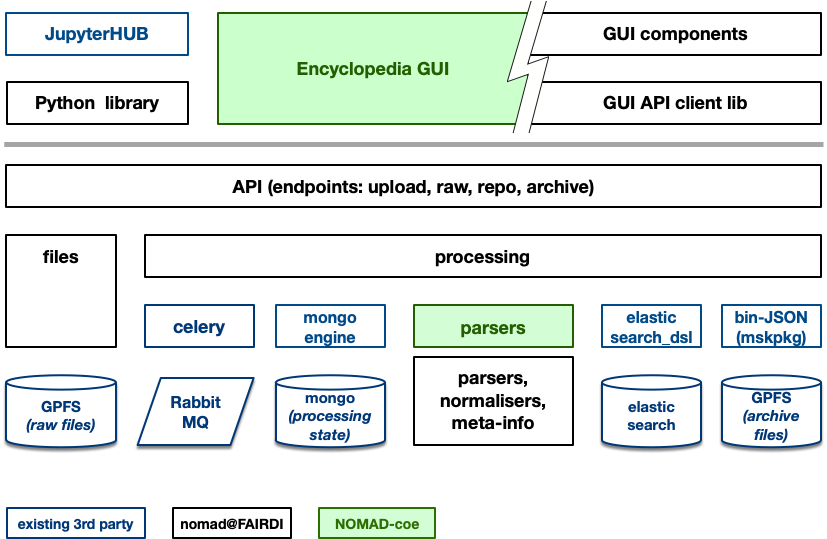
Besides various scientific computing, machine learning, and computational material science libraries (e.g. numpy, skikitlearn, tensorflow, ase, spglib, matid, and many more), Nomad uses a set of freely available or Open Source technologies that already solve most of its processing, storage, availability, and scaling goals. The following is a non comprehensive overview of used languages, libraries, frameworks, and services.
Python 3¶
The backend of nomad is written in Python. This includes all parsers, normalizers, and other data processing. We only use Python 3 and there is no compatibility with Python 2. Code is formatted close to pep8, critical parts use pep484 type-hints. Pycodestyle, pylint, and mypy (static type checker) are used to ensure quality. Tests are written with pytest. Logging is done with structlog and logstash (see Elasticstack below). Documentation is driven by Sphinx.
celery¶
Celery (+ rabbitmq) is a popular combination for realizing long running tasks in internet applications. We use it to drive the processing of uploaded files. It allows us to transparently distribute processing load while keeping processing state available to inform the user.
elastic search¶
Elasticsearch is used to store repository data (not the raw files). Elasticsearch allows for flexible scalable search and analytics.
mongodb¶
Mongodb is used to store and track the state of the processing of uploaded files and therein contained calculations. We use mongoengine to program with mongodb.
Keycloak¶
Keycloak is used for user management. It manages users and provide functions for registering, password forget, editing user accounts, and single sign on of fairdi@nomad and other related services.
flask, et al.¶
The ReSTful API is build with the flask framework and its ReST+ extension. This allows us to automatically derive a swagger description of the nomad API, which in turn allows us to generate programming language specific client libraries, e.g. we use bravado for Python and swagger-js for Javascript. Fruthermore, you can browse and use the API via swagger-ui.
Elasticstack¶
The elastic stack (previously ELK stack) is a central logging, metrics, and monitoring solution that collects data within the cluster and provides a flexible analytics frontend for said data.
Javascript, React, Material-UI¶
The frontend (GUI) of nomad@FAIRDI build on top of the React component framework. This allows us to build the GUI as a set of re-usable components to achieve a coherent representations for all aspects of nomad, while keeping development efforts manageable. React uses JSX (a ES6 variety) that allows to mix HTML with Javascript code. The component library Material-UI (based on Google’s popular material design framework) provides a consistent look-and-feel.
docker¶
To run a nomad@FAIRDI instance, many services have to be orchestrated: the nomad app, nomad worker, mongodb, Elasticsearch, Keycloak, RabbitMQ, Elasticstack (logging), the nomad GUI, and a reverse proxy to keep everything together. Further services might be needed (e.g. JypiterHUB), when nomad grows. The container platform Docker allows us to provide all services as pre-build images that can be run flexibly on all types of platforms, networks, and storage solutions. Docker-compose allows us to provide configuration to run the whole nomad stack on a single server node.
kubernetes + helm¶
To run and scale nomad on a cluster, you can use kubernetes to orchestrated the necessary containers. We provide a helm chart with all necessary service and deployment descriptors that allow you to setup and update nomad with few commands.
GitLab¶
Nomad as a software project is managed via GitLab. The nomad@FAIRDI project is hosted here. GitLab is used to manage versions, different branches of development, tasks and issues, as a registry for Docker images, and CI/CD platform.
Data model¶
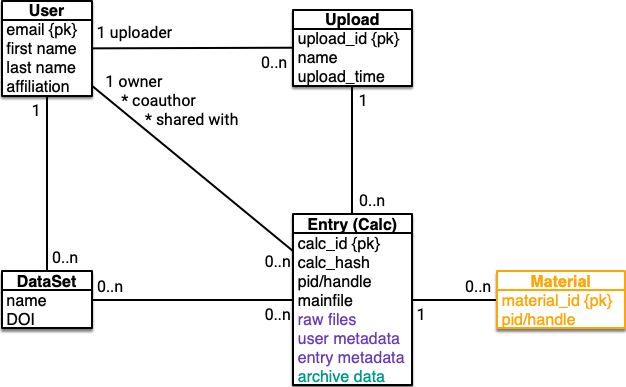
The entities that comprise the nomad data model are users, datasets, uploads, calculations (calc), and materials. Users upload multiple related calculations in one upload. Users can curate calculations into datasets. Caclulations belong to one material based on the simulated system.
Users¶
The user
emailis used as a primary key to uniquely identify users (even among different nomad installations)
Uploads¶
An upload contains related calculations in the form of raw code input and output files
Uploader are encouraged to upload all relevant files
The directory structure of an upload might be used to relate calculations to each other
Uploads have a unique randomly choosen
upload_id(UUID)The
uploaderis the user that provided the upload. There is always one immutableuploaderCurrently, uploads can be provided as
.zipor.tar.gzfiles.
Entries (Calculations, Code runs)¶
There are confusing names. Internally, in the nomad source code, the term
calcis used. An entry represents a single set of input/output used and produces by an individual run of a DFT code. If nomad is applied to other domains, i.e. experimental material science, entries might represent experiments or other entities.An entry (calculation) has a unique
calc_idthat is based on the upload’s id and themainfileThe
mainfileis a upload relative path to the main output file.Each calculation, when published, gets a unique
pid. Pids are ascending intergers. For eachpida shorterhandleis created. Handles can be registered with a handle system, e.g. the central nomad installation at MPCDF is registered at a MPCDF/GWDW handle system.The
calc_hashis computed from the main and other parsed raw files.Entry data comprises user metadata (comments, references, datasets, coauthors), calculation metadata (code, version, system and symmetry, used DFT method, etc.), the archive data (a hierarchy of all parsed quantities), and the uploaded raw files.
Datasets¶
Datasets are user curated sets of calculations.
Users can assign names and nomad can register a DOI for a dataset.
A calculation can be put into multiple datasets.
Materials¶
Materials aggregate calculations based on common system properties (e.g. system type, atoms, lattice, space group, etc.).
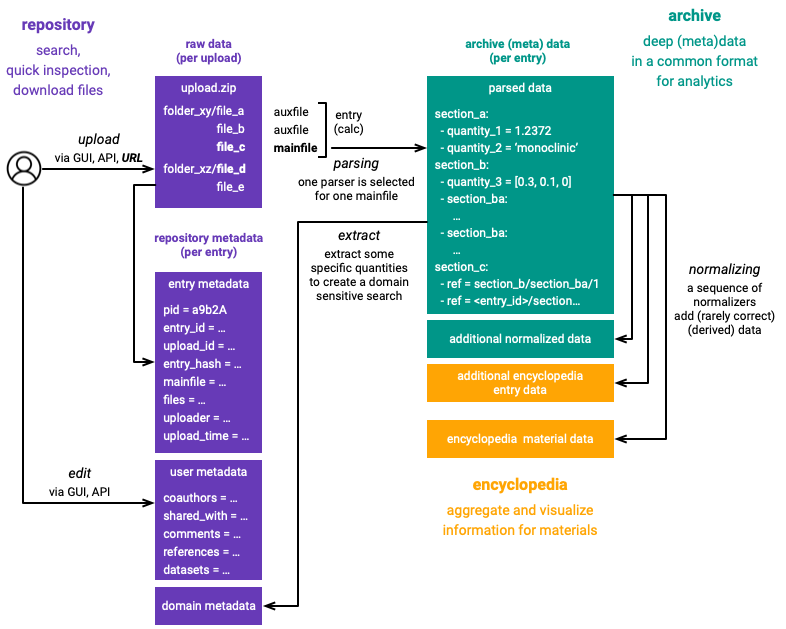
Data¶
We distinguish various forms of calculation data:
raw data: The raw files provided by nomad users
(repository) metadata: All data necessary to search and inspect nomad entries.
archive data: The data extracted from raw files by nomad parsers and normalizers. This data is represented in the meta-info format.
materials data: Aggregated information about calculations that simulated the same material.
Metadata¶
Metadata refers to those pieces of data, those quantities/attributes that we use to represent, identify, and index uploads and calculations in the API, search, GUI, etc. There are three catergories of metadata:
entry metadata: attributes that are necessary to uniquely identify entities (see also id-reference-label), that describe the upload, processing, etc. This data is derived by the nomad infrastructure.
user metadata: attributes provided by the user, e.g. comments, references, coauthors, datasets, etc. This data is provided by the user.
domain metadata: metadata parsed from raw files that describe calculations on a high level, e.g. code name, basis set, system type, etc. This data is derived from the uploaded data.
Those sets of metadata along with the actual raw and archive data are often transformed, passed, stored, etc. by the various nomad modules.
Implementation¶
The different entities have often multiple implementations for different storage systems.
For example, aspects of calculations are stored in files (raw files, calc metadata, archive data),
Elasticsearch (metadata), and mongodb (metadata, processing state).
Different transformation between different implementations exist. See
nomad.datamodel for further information.
Processing¶
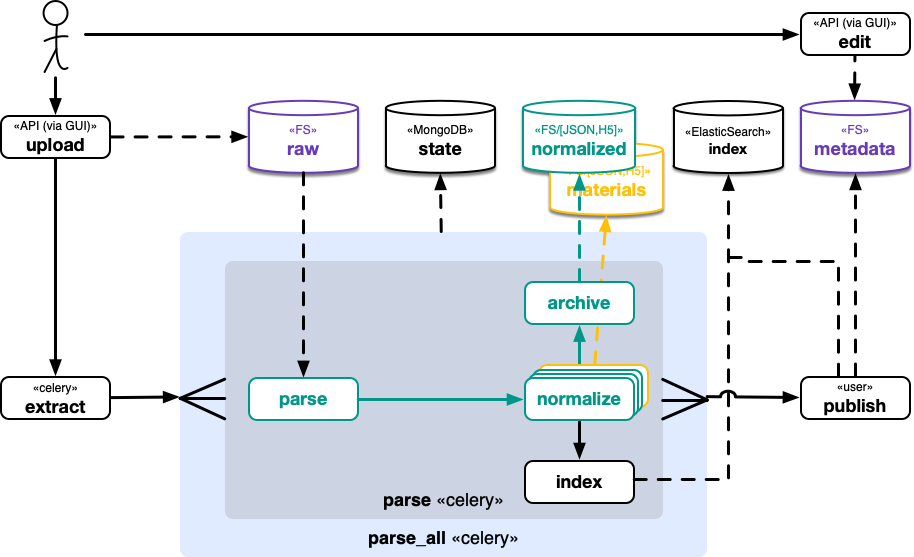
See nomad.processing for further information.